New method revolutionizes cancer diagnosis
Ground-breaking technology ‘Deep Visual Proteomics’ unites four technologies
How does cancer arise? How does cellular composition influence tumor malignancy? These questions are profound and challenging to answer, but are crucial to understand the disease and find the right cure. Now, a German-Danish team led by Professor Matthias Mann has developed a ground-breaking technology called ‘Deep Visual Proteomics’. This method provides researchers and clinicians with a protein read-out to understand cancer at single cell-type resolution. The technology was published in the journal Nature Biotechnology and demonstrates its potential in a first application to cancer cells.
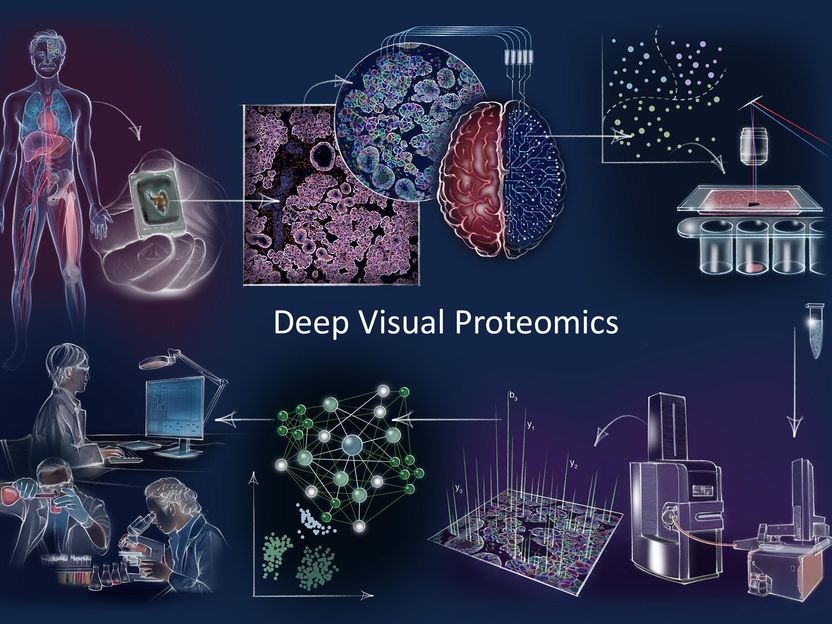
Deep Visual Proteomics concept and workflow Clockwise: Deep Visual Proteomics (DVP) combines high-resolution imaging, artificial intelligence (AI)-guided image analysis for single-cell classification and isolation with a novel ultra-sensitive proteomics workflow. DVP links data-rich imaging of cell culture or archived patient biobank tissues with deep learning-based cell segmentation and machine learning-based identification of cell types and states. (Un)supervised AI-classified cellular or subcellular objects of interest undergo automated laser microdissection and mass spectrometry (MS)-based proteomic profiling. Subsequent bioinformatics data analysis enables data mining to discover protein signatures providing molecular insights into proteome variation in health and disease states at the level of single cells.
© MPI of Biochemistry
Proteins are among the most important players in a variety of diseases. Aptly referred to as the 'molecular workhorses of the cell’, their proper function often determines the fitness of a cell and that of an individual by extension.
Matthias Mann explains: “When something goes wrong inside our cells and we become sick, you can be sure that proteins are involved in a wide range of different ways. Because of this, mapping the protein landscape can help us determine why a tumor could develop in a particular patient, what vulnerabilities that tumor has and also what treatment strategy might prove the most beneficial.”
Intrigued by these questions, a team of multi-disciplinary scientists from the research groups headed by Matthias Mann at the Max Planck Institute (MPI) of Biochemistry, near Munich and at the Novo Nordisk Foundation Center for Protein research (CPR) at University of Copenhagen have invented an innovative method called ‘Deep Visual Proteomics’. The study combines visual features of a tumor with a deep profiling technology to visualize the protein signatures in aberrant cells that are adjacent to the surrounding healthy cells. This would give the researchers an unprecedented insight into the disease and guide oncologists to develop clever strategies for diagnosis and treatment.
Deep Visual Proteomics unites four technologies
Deep Visual Proteomics integrates advances from four different technologies for the first time into a single workflow: First, advanced microscopy generates high-resolution tissue maps. Second, machine learning and artificial intelligence (AI) algorithms are used to accurately classify cells in shape, size, or protein localization before single cells are collected by highly-accurate laser capture microdissection. Then, after sorting normal or different types of diseased cells, the thousands of proteins present in the cell populations are detected all at once by using ultra-high sensitive mass spectrometry (MS) instruments from this minute amount of sample. Lastly, sophisticated bioinformatic analyses generate maps of proteins that provide spatial resolution of protein signatures across highly complex diseases such as cancer. Such protein landscapes are valuable tools for clinicians in understanding of mechanisms in health and disease in a greater detail.
“Our new concept, ‘Deep Visual Proteomics’, could become a game changer for molecular pathology in the hospitals. With this method, we take a tissue sample with tumor cells, and can identify and determine thousands of proteins in a minute of time and effort. These proteomic signatures reveal the mechanisms that drive tumor development and directly expose new therapeutic targets from a single tissue slice of a cancer patient biopsy. It exposes a cosmos of molecules inside these cancer cells,” says Andreas Mund, Associate Professor at CPR and part of Professor Matthias Mann’s team, that spearheaded this development.
Relevance for clinical pathology
In the new study, the researchers applied ‘Deep Visual Proteomics’ to cells from patients with acinic cell carcinoma and with melanoma. Lise Mette Rahbek Gjerdrum, consultant and clinical research associate professor at Department of Pathology, Zealand University Hospital and Department of Clinical Medicine, University of Copenhagen, describes: “This unique method combines tissue architecture with the expression of thousands of proteins specific for selected cells. It enables researchers to investigate interactions between cancer cells and their surrounding cells with major implications for future clinical cancer treatment. Recently, we diagnosed a highly complex clinical case using ‘Deep Visual Proteomics’.’’
Dr. Fabian Coscia, co-first author of the study and – since June 2021 – head of the research group the "Spatial Proteomics" at the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) in Berlin, says: "But of course, the technique can be used to characterize all other tumor types in similar details." His goal, he says, is to tap into the archived patient biobanks and find new targets for individualized, i.e., patient-tailored cancer therapies - even for previously treatment-resistant tumors.
And it is not only cancer that can be better understood using ‘Deep Visual Proteomics’. The method can also be applied to other diseases. "For example, you can analyze the proteins in nerve cells to find out exactly what happens in the cells in neurodegenerative diseases such as Alzheimer's or Parkinson's." says Fabian Coscia further.
‘’By merging the power of microscopy, AI and ultra-sensitive MS-based proteomics, we have developed a method that is very powerful to understand the molecular wiring of healthy versus disease cells. This could help doctors identify targets for future drugs and diagnostics.’’ concludes Matthias Mann.
Original publication
A. Mund*#, F. Coscia*, A. Kriston, R. Hollandi, F. Kovács, A.-D. Brunner, E. Migh, L. Schweizer, A. Santos Delgado, M. Bzorek, S. Naimy, L.M. Rahbek Gjerdrum7,13, B. Dyring-Andersen, J. Bulkescher, C. Lukas, M.A. Eckert, E. Lengyel, C. Gnann, E. Lundberg, P. Horvath#, M. Mann#; * shared first-authors and # corresponding authors; Deep Visual Proteomics defines cell identity and heterogeneity; Nature Biotechnology, May 2022
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Mass Spectrometry
Mass spectrometry enables us to detect and identify molecules and reveal their structure. Whether in chemistry, biochemistry or forensics - mass spectrometry opens up unexpected insights into the composition of our world. Immerse yourself in the fascinating world of mass spectrometry!
Topic World Mass Spectrometry
Mass spectrometry enables us to detect and identify molecules and reveal their structure. Whether in chemistry, biochemistry or forensics - mass spectrometry opens up unexpected insights into the composition of our world. Immerse yourself in the fascinating world of mass spectrometry!