Breaking a chemical bond
A new theory for the breaking of (bio-)chemical bonds under load may help to predict the strength and performance of synthetic nanostructures and proteins, on a molecular level.
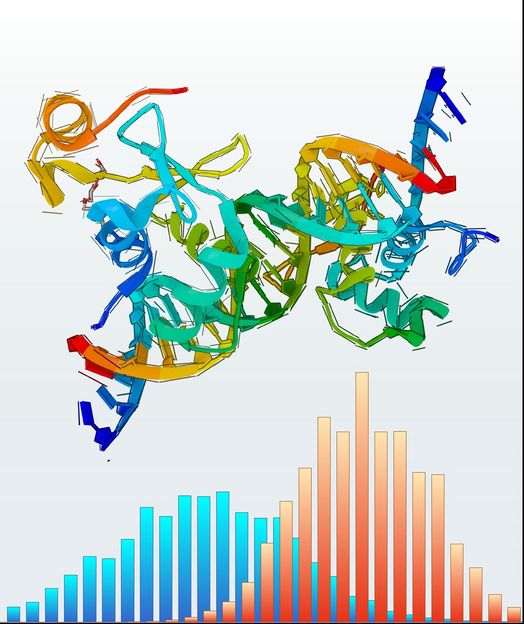
The illustration schematically depicts a protein structure and rupture force distributions of one of its macromolecular bonds under load for two different loading rates.
Klaus Kroy
The fundamental question how a molecular bond breaks is of interest in many fields of science and has been studied extensively. Yet, now writing in Nature Communications, a group of theoretical physicists from the University of Leipzig, Germany, has put forward a more powerful analytical formula for forcible bond breaking than previously available. It predicts how likely a bond will break at a given load, if probed with a prescribed loading protocol. This so-called rupture force distribution is the most informative and most commonly measured quantity in modern single-molecule force spectroscopy experiments (which may roughly be thought of as nanoscopic versions of the conventional crash- or breaking tests employed in materials science and engineering). Such experiments are nowadays performed in large numbers in molecular biology and biophysics labs to probe the mechanical strength of individual macromolecular bonds.
Recent methodological advances have pushed force spectroscopy assays to ever higher loading rates (the equivalent of the speed employed in the macroscopic crash-test). This provided a strong incentive for the Leipzig team to improve on current state-of-the-art theories for forcible bond breaking, which are limited to comparatively low speeds. Moreover, the new equation solves another problem that has bothered experts in the field for many years. Force spectroscopy experiments are often simulated with sophisticated all-atom computer models to supplement the experimental data with information on internal molecular details that cannot be resolved in a laboratory setting. However, because of their enormous complexity, such computer simulations operate at extremely high loading rates to cut down on the runtime. As a consequence, simulation and experiment were so far two essentially distinct branches of force spectroscopy. The new equation, which gives exact results for both low and high loading rates, will thus suit both experimentalists and computer scientists, and help them to systematically analyze and compare their results. This should eventually improve our microscopic understanding of the strength of synthetic materials and of how proteins attain and maintain their three-dimensional structure and perform conformational changes, which are core features determining the function and dysfunction of these
amazing engines of life.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Spectroscopy
Investigation with spectroscopy gives us unique insights into the composition and structure of materials. From UV-Vis spectroscopy to infrared and Raman spectroscopy to fluorescence and atomic absorption spectroscopy, spectroscopy offers us a wide range of analytical techniques to precisely characterize substances. Immerse yourself in the fascinating world of spectroscopy!

Topic World Spectroscopy
Investigation with spectroscopy gives us unique insights into the composition and structure of materials. From UV-Vis spectroscopy to infrared and Raman spectroscopy to fluorescence and atomic absorption spectroscopy, spectroscopy offers us a wide range of analytical techniques to precisely characterize substances. Immerse yourself in the fascinating world of spectroscopy!




























































