New software processes huge amounts of single-cell data
Comprehensive analysis of large gene-expression datasets
Scientists from the Helmholtz Zentrum München have developed a program that is able to help manage enormous datasets. The software, named Scanpy, is a candidate for analyzing the Human Cell Atlas.
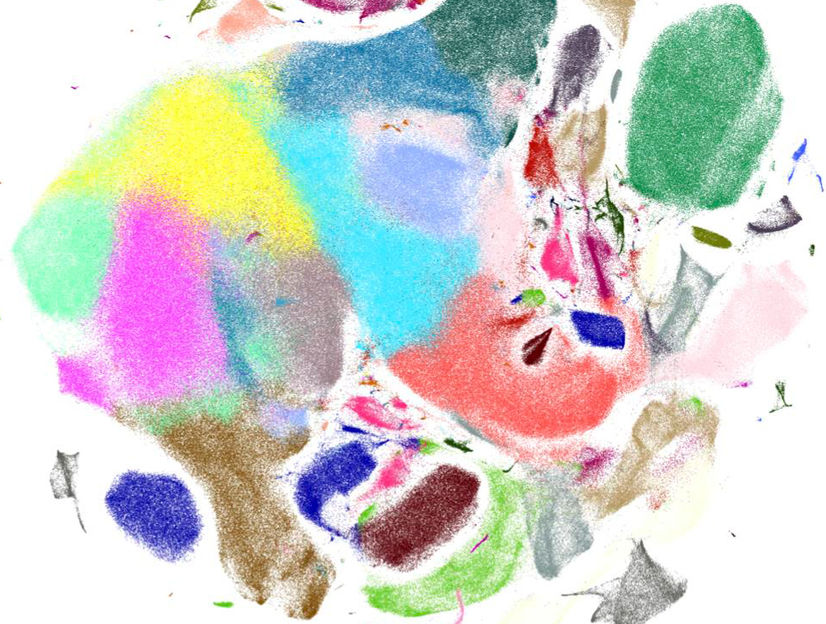
Visualization of gene expression patterns of murine brain cells generated with Scanpy.
Helmholtz Zentrum München
“It’s about analyzing gene-expression data of a large number of individual cells,” explains lead author Alex Wolf of the Institute of Computational Biology (ICB) at Helmholtz Zentrum München. He developed Scanpy together with his colleague Philipp Angerer in the Machine Learning Group of Prof. Dr. Dr. Fabian Theis. In addition to his position at Helmholtz Zentrum, Theis is also a professor of mathematical modelling of biological systems at the Technical University of Munich. “New technical advances generate several orders of magnitude more data with a correspondingly greater information content,” Theis says. “However, the historically evolved software infrastructure for gene-expression analysis simply wasn’t designed to cope with the new challenges. New analytic methods are therefore needed.”
The race for the Human Cell Atlas
According to Theis, a major international research project could also benefit from the software. A team of international scientists is compiling a reference database, called the Human Cell Atlas, which holds data on the gene activity of all human cell types. “For this project, and in a growing number of other projects in which databases are combined, it is important to have scalable software,” says Theis. It is therefore no surprise that Scanpy is currently a candidate for helping to analyze the Human Cell Atlas.
“The publication of Scanpy marks the first software that allows comprehensive analysis of large gene-expression datasets with a broad range of machine-learning and statistical methods,” explains Wolf, describing the achievement. “The software is already being used by a number of groups around the world, notably at the Broad Institute of Harvard University and the Massachusetts Institute of Technology, MIT.”
Technologically, the application is a trailblazing development: Whereas biostatistics programs are traditionally written in the programming language R, Scanpy is based on the Python language, the dominant language in the machine learning community. Another new feature is that graph-based algorithms lie at the heart of Scanpy. Unlike the usual approach of regarding cells as points in a coordinate system within gene-expression space, the algorithms use a graph-like coordinate system. Instead of characterizing a single cell by the expression value for thousands of genes, the system simply characterizes cells by identifying their closest neighbors – very much like the connections in social networks. In fact, to identify cell types, Scanpy uses the same algorithms as Facebook does for identifying communities.
Original publication
Other news from the department science
These products might interest you

Get the analytics and lab tech industry in your inbox
By submitting this form you agree that LUMITOS AG will send you the newsletter(s) selected above by email. Your data will not be passed on to third parties. Your data will be stored and processed in accordance with our data protection regulations. LUMITOS may contact you by email for the purpose of advertising or market and opinion surveys. You can revoke your consent at any time without giving reasons to LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin, Germany or by e-mail at revoke@lumitos.com with effect for the future. In addition, each email contains a link to unsubscribe from the corresponding newsletter.