DNA from extinct humans discovered in cave sediments
New method retrieve hominin DNA from cave sediments – even in the absence of skeletal remains
While there are numerous prehistoric sites in Europe and Asia that contain tools and other human-made artefacts, skeletal remains of ancient humans are scarce. Researchers of the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, have therefore looked into new ways to get hold of ancient human DNA. From sediment samples collected at seven archaeological sites, the researchers “fished out” tiny DNA fragments that had once belonged to a variety of mammals, including our extinct human relatives. They retrieved DNA from Neandertals in cave sediments of four archaeological sites, also in layers where no hominin skeletal remains have been discovered. In addition, they found Denisovan DNA in sediments from Denisova Cave in Russia. These new developments now enable researchers to uncover the genetic affiliations of the former inhabitants of many archaeological sites which do not yield human remains.
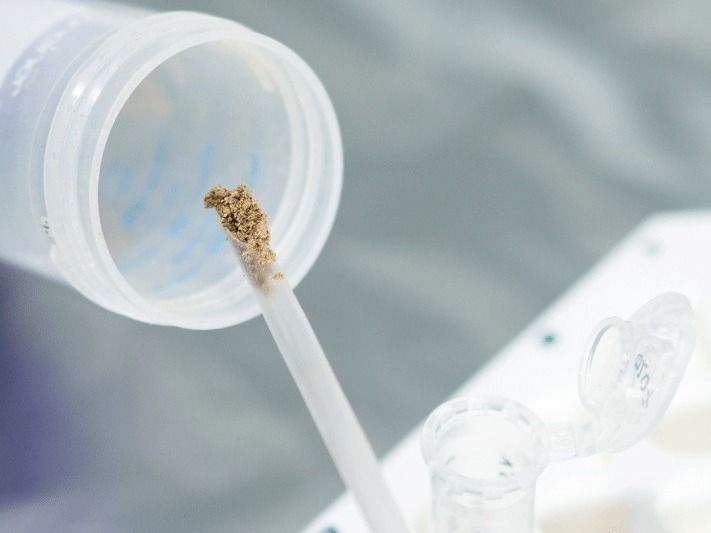
A sediment sample is prepared for DNA extraction.
© MPI f. evolutionäre Anthropologie/ S. Tüpke
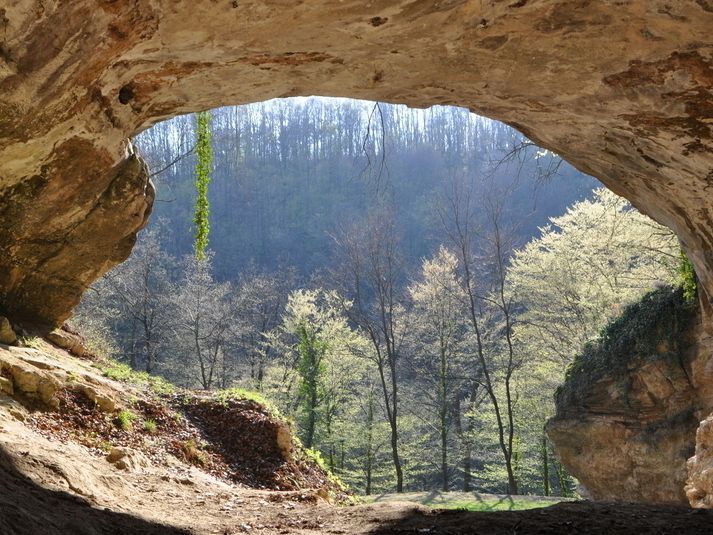

Entrance to the archaeological site of Vindija Cave, Croatia.
© MPI f. evolutionäre Anthropologie/ J. Krause

By looking into the genetic composition of our extinct relatives, the Neandertals, and their cousins from Asia, the Denisovans, researchers can shed light on our own evolutionary history. However, fossils of ancient humans are rare, and they are not always available or suitable for genetic analyses. “We know that several components of sediments can bind DNA”, says Matthias Meyer of the Max Planck Institute for Evolutionary Anthropology. “We therefore decided to investigate whether hominin DNA may survive in sediments at archaeological sites known to have been occupied by ancient hominins.”
To this aim Meyer and his team collaborated with a large network of researchers excavating at seven archaeological sites in Belgium, Croatia, France, Russia and Spain. Overall, they collected sediment samples covering a time span from 14,000 to over 550,000 years ago. Using tiny amounts of material the researchers recovered and analyzed fragments of mitochondrial DNA – genetic material from the mitochondria, the “energy factories” of the cell – and identified them as belonging to twelve different mammalian families that include extinct species such as the woolly mammoth, the woolly rhinoceros, the cave bear and the cave hyena.
The researchers then looked specifically for ancient hominin DNA in the samples. “From the preliminary results, we suspected that in most of our samples, DNA from other mammals was too abundant to detect small traces of human DNA”, says Viviane Slon, Ph.D. student at the Max Planck Institute in Leipzig and first author of the study. “We then switched strategies and started targeting specifically DNA fragments of human origin.” Nine samples from four archaeological sites contained enough ancient hominin DNA for further analyses: Eight sediment samples contained Neandertal mitochondrial DNA from either one or multiple individuals, while one sample contained Denisovan DNA. Most of these samples originated from archaeological layers or sites where no Neandertal bones or teeth were previously found.
A new tool for archaeology
“By retrieving hominin DNA from sediments, we can detect the presence of hominin groups at sites and in areas where this cannot be achieved with other methods”, says Svante Pääbo, director of the Evolutionary Genetics department at the Max Planck Institute for Evolutionary Anthropology and co-author of the study. “This shows that DNA analyses of sediments are a very useful archaeological procedure, which may become routine in the future”.
Even sediment samples that were stored at room temperature for years still yielded DNA. Analyses of these and of freshly-excavated sediment samples recovered from archaeological sites where no human remains are found will shed light on these sites’ former occupants and our joint genetic history.
Original publication
Viviane Slon et al.; "Neandertal and Denisovan DNA from Pleistocene sediments"; Science; 27 April, 2017























































