Real-time observation of enzymatic processes on DNA
Insights into the molecular processes involved in the detection of DNA strand breaks
DNA strand breaks can contribute to the development of cancer and the ageing process. Researchers from the Departments of Biology and Chemistry of the University of Konstanz have now been able to observe in real time the molecular processes that take place at DNA strand breaks by means of infrared spectroscopy.
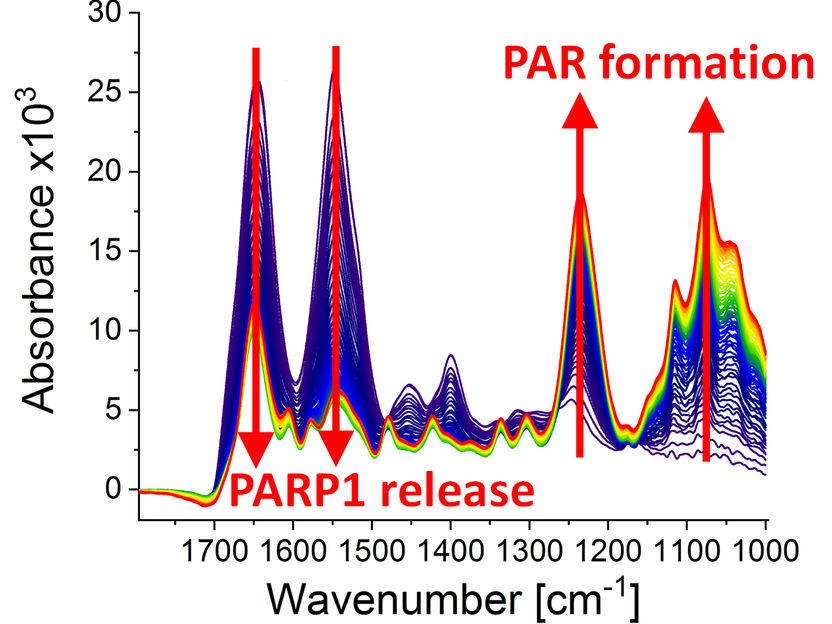
Infrared spectra at different points of time (0-79 min) after the poly(ADP-ribosyl)ation reaction started due to the addition of PARP1 substrate NAD+. The following can be observed: the dynamic formation of the biopolymer poly(ADP-ribose) (absorption bands at 1236 cm-1 and 1074 cm-1) and the detachment of PARP1 from the DNA strand break (absorption bands at 1645 cm-1 and 1548 cm-1).
Modified from Krüger et al., “Real-time monitoring of PARP1-dependent PARylation by ATR-FTIR-spectroscopy”, Nature Communications, 1 May 2020. DOI: 10.1038/s41467-020-15858-w.
DNA damage in general and DNA strand breaks in particular occur every day in all cells of the human body. This is due to internal influences such as free radicals, which are produced during inflammatory processes and cellular respiration, and external ones, such as cosmic background radiation or X-rays in the course of medical diagnostic measures. DNA strand breaks can lead to cell death or to mutations and thus contribute in the long term to cancer development or the ageing process.
DNA repair by PARP1
Cells possess molecular tools to repair such DNA strand breaks very efficiently. One of them is the enzyme poly(ADP-ribose) polymerase 1 (PARP1), which detects DNA strand breaks and thereby initiates downstream repair processes. By binding to a DNA strand break, PARP1 is (catalytically) activated and uses the substrate nicotinamide adenine dinucleotide (NAD+) to produce poly(ADP-ribose) (PAR), a chain-shaped biopolymer. This serves as a signal transmitter in the cell and coordinates the further DNA damage response. In the further course of the process, PARP1 detaches from the site of damage again, thus clearing the way for subsequent steps in DNA repair. This process is of medical importance, even more so as pharmacological inhibitors of PARP1 have recently been introduced into cancer therapy.
Scientists at the University of Konstanz (working groups of Professor Aswin Mangerich and Professor Alexander Bürkle, Department of Biology, and working group of Professor Karin Hauser, Department of Chemistry) have now been able to visualize in detail the biochemical processes which PARP1 fulfills at a DNA strand break. To this end, they used a special method of infrared spectroscopy (ATR-FTIR), which had also been successfully used in a previous, recently published study on the interactions of the tumour suppressor protein p53 with DNA and PAR.
Real-time observations using infrared spectroscopy
“What is special about our new study is that we can now investigate the molecular processes that PARP1 undergoes at DNA strand breaks in real time. This enabled us to unveil dynamic changes in the protein structure and thus gain further insights into the underlying mechanisms”, said Dr Annika Krüger, who worked on the project as part of her – by now successfully completed – doctoral thesis. During the work on her doctoral thesis, Krüger was supported by the Research School Chemical Biology, the Zukunftskolleg of the University of Konstanz and the Konstanz Collaborative Research Centre 969 “Chemical and Biological Principles of Cellular Proteostasis”. She is now pursuing research at the renowned Karolinska Institute in Stockholm, Sweden.
As a matter of principle, this spectroscopic method can be used to investigate also other enzymatic processes that take place at the DNA, in detail and with molecular resolution. In the long term, this may contribute to better understanding of the mechanisms of cancer development and ageing, as well as of the mode of action of anticancer drugs.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Spectroscopy
Investigation with spectroscopy gives us unique insights into the composition and structure of materials. From UV-Vis spectroscopy to infrared and Raman spectroscopy to fluorescence and atomic absorption spectroscopy, spectroscopy offers us a wide range of analytical techniques to precisely characterize substances. Immerse yourself in the fascinating world of spectroscopy!

Topic World Spectroscopy
Investigation with spectroscopy gives us unique insights into the composition and structure of materials. From UV-Vis spectroscopy to infrared and Raman spectroscopy to fluorescence and atomic absorption spectroscopy, spectroscopy offers us a wide range of analytical techniques to precisely characterize substances. Immerse yourself in the fascinating world of spectroscopy!























































