Using quantum dots and a smartphone to find killer bacteria
Scientists develop cheap and rapid way to identify antibiotic-resistant golden staph (MRSA)
A combination of off-the-shelf quantum dot nanotechnology and a smartphone camera soon could allow doctors to identify antibiotic-resistant bacteria in just 40 minutes, potentially saving patient lives.
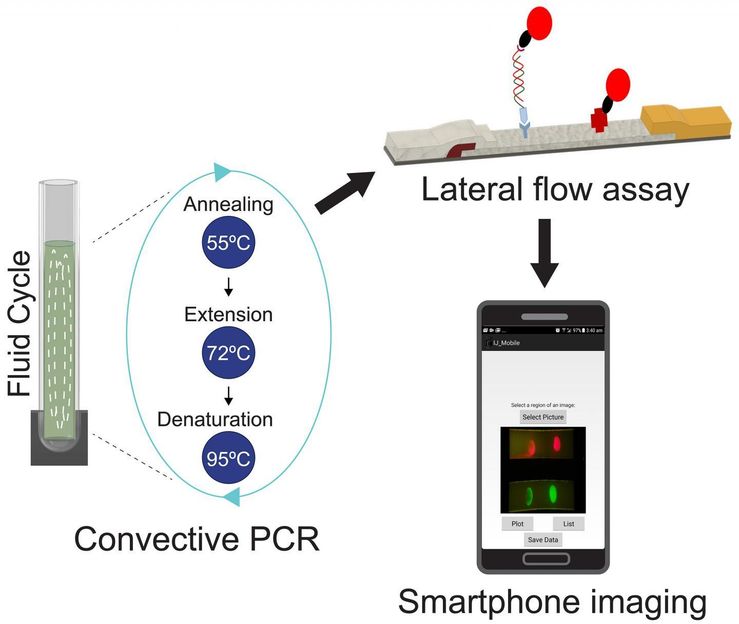
This is a diagram depicting the analysis process.
Sunna Lab
Staphylococcus aureus (golden staph), is a common form of bacterium that causes serious and sometimes fatal conditions such as pneumonia and heart valve infections. Of particular concern is a strain that does not respond to methicillin, the antibiotic of first resort, and is known as methicillin-resistant S. aureus, or MRSA.
Recent reports estimate that 700000 deaths globally could be attributed to antimicrobial resistance, such as methicillin-resistance. Rapid identification of MRSA is essential for effective treatment, but current methods make it a challenging process, even within well-equipped hospitals.
Soon, however, that may change, using nothing except existing technology.
Researchers from Macquarie University and the University of New South Wales, both in Australia, have demonstrated a proof-of-concept device that uses bacterial DNA to identify the presence of Staphylococcus aureus positively in a patient sample - and to determine if it will respond to frontline antibiotics.
In a paper published in the international peer-reviewed journal Sensors and Actuators B: Chemical the Macquarie University team of Dr Vinoth Kumar Rajendran, Professor Peter Bergquist and Associate Professor Anwar Sunna with Dr Padmavathy Bakthavathsalam (UNSW) reveal a new way to confirm the presence of the bacterium, using a mobile phone and some ultra-tiny semiconductor particles known as quantum dots.
"Our team is using Synthetic Biology and NanoBiotechnology to address biomedical challenges. Rapid and simple ways of identifying the cause of infections and starting appropriate treatments are critical for treating patients effectively," says Associate Professor Anwar Sunna, head of the Sunna Lab at Macquarie University.
"This is true in routine clinical situations, but also in the emerging field of personalised medicine."
The researchers' approach identifies the specific strain of golden staph by using a method called convective polymerase chain reaction (or cPCR). This is a derivative of a widely -employed technique in which a small segment of DNA is copied thousands of times, creating multiple samples suitable for testing.
Vinoth Kumar and colleagues then subject the DNA copies to a process known as lateral flow immunoassay - a paper-based diagnostic tool used to confirm the presence or absence of a target biomarker. The researchers use probes fitted with quantum dots to detect two unique genes, that confirms the presence of methicillin resistance in golden staph
A chemical added at the PCR stage to the DNA tested makes the sample fluoresce when the genes are detected by the quantum dots - a reaction that can be captured easily using the camera on a mobile phone.
The result is a simple and rapid method of detecting the presence of the bacterium, while simultaneously ruling first-line treatment in or out.
Although currently at proof-of-concept stage, the researchers say their system which is powered by a simple battery is suitable for rapid detection in different settings.
"We can see this being used easily not only in hospitals, but also in GP clinics and at patient bedsides," says lead author, Macquarie's Vinoth Kumar Rajendran.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World PCR
This groundbreaking and highly versatile molecular technique of PCR allows us to amplify tiny amounts of genetic material on a large scale and analyze them in detail. Whether in medical diagnostics, forensic DNA analysis or research into genetic diseases - PCR is an indispensable tool that gives us deep insights into the world of DNA. Immerse yourself in the fascinating world of the polymerase chain reaction (PCR)!

Topic World PCR
This groundbreaking and highly versatile molecular technique of PCR allows us to amplify tiny amounts of genetic material on a large scale and analyze them in detail. Whether in medical diagnostics, forensic DNA analysis or research into genetic diseases - PCR is an indispensable tool that gives us deep insights into the world of DNA. Immerse yourself in the fascinating world of the polymerase chain reaction (PCR)!

























































