Proteins may prevent dysfunction and disease by relaxing
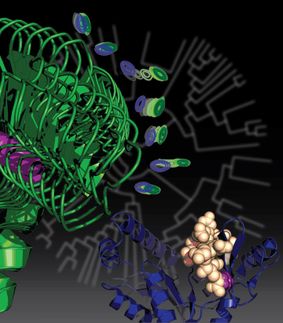
For many years, we thought that all proteins must fold into complicated shapes to fulfill their functions, looking like thousands of sets of custom-tailored locks and keys. But over the past two decades, scientists have begun to realize other proteins -- including those involved in many essential cellular functions -- remain fully or partially unfolded for parts of their lives.
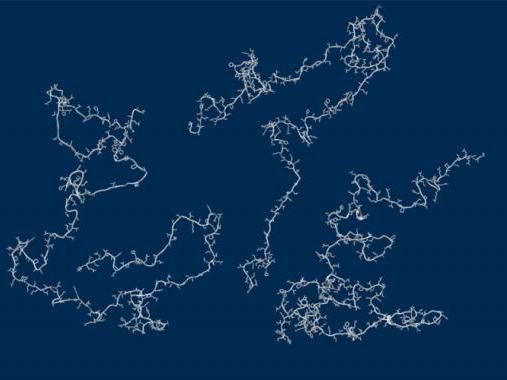

A new study suggests many proteins remain expanded in the cell, rather than contracting into tight folded shapes.
Micayla Bowman
Out of this realization has come a debate: how such proteins spend their time when floating in the cell. Since they're not folded into their precise shapes, do they contract into balls, or remain expanded, like a rope? Different methods have yielded different results. Scattering using X-rays suggested they remained expanded, but fluorescent methods observed the opposite behavior. The answer would affect how we envision the movement of a protein through its life -- essential for understanding how proteins fold, what goes wrong during disorders and disease and how to model their behavior.
A team of University of Chicago and Notre Dame researchers used simulations and X-rays to tackle the question. The results conclude that these proteins remain unfolded and expanded as they float loose in the cytoplasm of a cell.
"What we need to determine is how 'sticky' they are," said co-author Prof. Tobin Sosnick, chair of the Department of Biochemistry & Molecular Biology, director of the Graduate Program in Biophysical Sciences and member of both the Computation Institute and the Institute for Molecular Engineering. "For survival, you want a protein to "stick" only in the right conformation, and with the right other pieces or partners at the right times." This stickiness is determined by the protein's chemistry and physics.
Sosnick's team came up with a way to analyze a single X-ray measurement to determine stickiness, finding that the proteins are generally much less sticky than expected. Their chemistry is set up prefer to interact with water, rather than itself or other proteins.
"The results are very clear, but contradict the current consensus," Sosnick said. "It's possible that proteins can avoid unwanted interactions by being expanded." This reduces the chances of them sticking to other proteins by accident, causing dysfunction or disease.
Original publication
Riback, Joshua A. and Bowman, Micayla A. and Zmyslowski, Adam M. and Knoverek, Catherine R. and Jumper, John M. and Hinshaw, James R. and Kaye, Emily B. and Freed, Karl F. and Clark, Patricia L. and Sosnick, Tobin R.; "Innovative scattering analysis shows that hydrophobic disordered proteins are expanded in water"; Science; 2017
Original publication
Riback, Joshua A. and Bowman, Micayla A. and Zmyslowski, Adam M. and Knoverek, Catherine R. and Jumper, John M. and Hinshaw, James R. and Kaye, Emily B. and Freed, Karl F. and Clark, Patricia L. and Sosnick, Tobin R.; "Innovative scattering analysis shows that hydrophobic disordered proteins are expanded in water"; Science; 2017
Organizations
Other news from the department science

Get the analytics and lab tech industry in your inbox
From now on, don't miss a thing: Our newsletter for analytics and lab technology brings you up to date every Tuesday. The latest industry news, product highlights and innovations - compact and easy to understand in your inbox. Researched by us so you don't have to.