Photonic-plasmonic microcavity for ultrasensitive protein detection
Measuring proteins in real-time down to fM solution concentration levels
Label free optical biosensors enable the monitoring of biomolecules and their interactions in often highly sensitive diagnostic assays. Several methods have been employed for this purpose, including Whispering Gallery Mode (WGM) biosensing, which offers a particularly sensitive approach to quantify the mass loading of biomolecules on the resonator surface with ultimate sensitivity estimated on the single molecule level. The simplest WGM biosensor is a glass microsphere (typically 50–100 mm in diameter) where the resonant light remains confined by total-internal reflection.
WGM sensors derive their unprecedented sensitivity from the use of high quality-factor (Q-factor) optical resonances to monitor wavelength shift signals upon binding of biomolecules or nanobeads to the resonator surface. Even a single virus could be detected. Yet, if e.g. a single protein molecule shall be detected, the sensitivity has to be boosted. There have been several approaches, such as the generation of hot spots using a hybrid photonic-plasmonic sensing concept with a gold nanoparticle (NP) layer coupled to a WGM biosensor. However, there are some drawbacks: First, measurements cannot be done directly in solution. Second, real-time analysis is not possible since the proteins have to be pre-adsorbed on the NPs. Third, proteins are adsorbed randomly within the NP layer – outside of plasmonic field enhancements sites – which lowers the detection sensitivity.
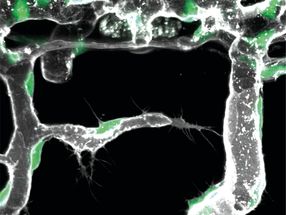
A German-American team led by Frank Vollmer and Melik C. Demirel now proposes an alternative concept overcoming these problems: optical trapping of protein molecules at the sites of plasmonic field enhancements in a random gold NP layer. The stable integration of the microsphere WGM biosensor with a wetted gold NP layer is critical for achieving ultra-sensitive detection. Therefore, the silica microsphere cavity remains fixed on the Au NP layer. The Q-factor of the microsphere drops slightly but is still in the 105 range. After adding bovine serum albumin (BSA) solution at microliter of sample volumes, which enters the NP layer by capillary suction, the researchers observed an unexpectedly large significant wavelength shift.
The achieved sensitivity in the order of femtomole concentration levels was very surprising, and cannot be explained from random binding of the BSA molecules to the NP surface. Instead, the scientists hypothesized that the protein molecules prefer to bind to hotspot locations (i.e. closely spaced random NPs) of plasmon resonances excited in the NP layer due to optical trapping. To validate this hypothesis, they calculated the electromagnetic field distribution in a model NP layer using generalized Mie theory and simulated the expected wavelength shift due to the binding of proteins. Their calculations showed that, indeed, optical trapping of the proteins at highly sensitive plasmonic hotspot locations is essential for achieving high sensitivity in microcavity biosensing.
The achieved sensitivity in the order of femtomole concentration levels was very surprising, and cannot be explained from random binding of the BSA molecules to the NP surface. Instead, the scientists hypothesized that the protein molecules prefer to bind to hotspot locations (i.e. closely spaced random NPs) of plasmon resonances excited in the NP layer due to optical trapping. To validate this hypothesis, they calculated the electromagnetic field distribution in a model NP layer using generalized Mie theory and simulated the expected wavelength shift due to the binding of proteins. Their calculations showed that, indeed, optical trapping of the proteins at highly sensitive plasmonic hotspot locations is essential for achieving high sensitivity in microcavity biosensing.
Original publication
Santiago-Cordoba, M. A., et al.; J. Biophotonics 5(8-9), 629-638 (2012)
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic world Diagnostics
Diagnostics is at the heart of modern medicine and forms a crucial interface between research and patient care in the biotech and pharmaceutical industries. It not only enables early detection and monitoring of disease, but also plays a central role in individualized medicine by enabling targeted therapies based on an individual's genetic and molecular signature.

Topic world Diagnostics
Diagnostics is at the heart of modern medicine and forms a crucial interface between research and patient care in the biotech and pharmaceutical industries. It not only enables early detection and monitoring of disease, but also plays a central role in individualized medicine by enabling targeted therapies based on an individual's genetic and molecular signature.